Trainable Models
The ‘Trainable Model’ concept utilizing a novel similarity based analysis methodology allows the user to:
- Assess the quality of the predictions by means of the Reliability Index (RI) estimation. This index provides values in a range from 0 to 1 and serves as an evaluation of whether a submitted compound falls within the Model Applicability Domain. Estimation of the Reliability Index takes into account the following two aspects: similarity of the tested compound to the training set and the consistency of experimental values for similar compounds.
- Instantly expand the Model Applicability Domain with the help of any user-defined proprietary ‘in-house’ data of experimental values for the property of interest.
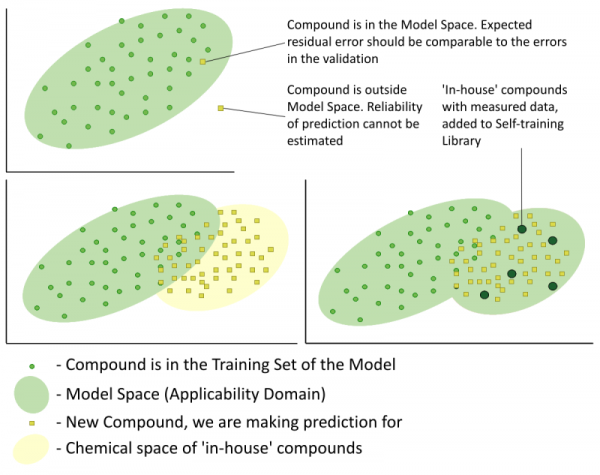
Each ‘Trainable Model’ consists of the following parts:
- A structure based QSAR/QSPR for the prediction of the property of interest derived from a literature training set – the baseline QSAR/QSPR.
- A user defined data set with experimental values for the property of interest – the Self-training Library.
- A special similarity based routine which identifies the most similar compounds contained in the Self-training Library and considering their experimental values calculates systematic deviations produced by the baseline QSAR/QSPR for each submitted molecule – the training engine.
Note: The Reliability Index (RI) is calculated as a product of two underlying factors, both ranging from 0 to 1: Similarity Index (SI) reflecting the overall similarity to the compounds in the training library, and Data-model Consistency Index (DCI) indicating how consistent is the performance of the baseline model for the most similar compounds in the library. SI and DCI values are not displayed in Prediction Modules, but can optionally be included in the output of calculations in Spreadsheet (see option 6g here). For more technical details of how these parameters are calculated one can refer to [1].
The current version of ACD/Percepta has implemented ‘Trainable Model’ methodology for the prediction of the following properties:
- P-gp Specificity
- Trainable P-gpS
Calculates the probability of a compound being a P-gp substrate. - Trainable P-gpI
Predicts the probability for a compound to act as a P-gp inhibitor.
- Trainable P-gpS
- Solubility
- Trainable LogS0
Calculates intrinsic solubility in water (LogS0, mmol/ml). - Trainable LogS
Calculates solubility in buffer at relevant pH values (LogS, mmol/ml).
Training takes place through LogS0 Self-training Libraries.
- Trainable LogS0
- Plasma Protein Binding
- Trainable LogKa(HSA)
Predicts the compound's equilibrium binding constant to human serum albumin in the blood plasma (LogKaHSA). - Trainable PPB
Estimates the fraction of the compound bound to the blood plasma proteins (%PPB)
- Trainable LogKa(HSA)
- Partitioning
- Trainable LogP
Calculates the logarithm of the octanol-water partitioning coefficient for the neutral form of the compound (LogP) - Trainable LogD
Calculates the logarithm of the apparent octanol water partition coefficient at relevant pH values (LogD) taking into account all the species (including ionized) of the compound present in the solution.
Training takes place through LogP Self-training Libraries.
- Trainable LogP
- Cytochrome P450 Inhibitor Specificity
Calculates probability of a compound being an inhibitor of a particular cytochrome P450 enzyme with IC50 below one of the two selected thresholds (general inhibition models - IC50 < 50 μM; efficient inhibition - IC50 < 10 μM). Predictions are available for five P450 isoforms :- Trainable CYP1A2 I
- Trainable CYP2C19 I
- Trainable CYP2C9 I
- Trainable CYP2D6 I
- Trainable CYP3A4 I
- Cytochrome P450 Substrate Specificity
Calculates probability of a compound being metabolized by a particular cytochrome P450 enzyme. Predictions are available for five P450 isoforms:- Trainable CYP1A2 S
- Trainable CYP2C19 S
- Trainable CYP2C9 S
- Trainable CYP2D6 S
- Trainable CYP3A4 S
- Solubility in DMSO
- Trainable DMSO Solubility
Calculates probability of the compound's solubility in DMSO exceeding 20 mM threshold.
- Trainable DMSO Solubility
- Aquatic Toxicity
- Trainable LC50 D. magna
Calculates the median lethal concentration (LC50) of a compound to crustacean species Daphnia magna. - Trainable LC50 P. promelas
Calculates the median lethal concentration (LC50) of a compound to fish species Pimephales promelas. - Trainable IGC50 T. pyriformis
Calculates the median inhibition growth concentration (IGC50) of a compound to protozoan species Tetrahymena pyriformis.
- Trainable LC50 D. magna
- Acute Toxicity
Calculates the median lethal dose (LD50) of a compound for different species (mouse and rat) and administered by different routes (intraperitoneal, intravenous, oral, subcutaneous). Predictions are available for six species/administration route combinations:- Trainable LD50 Mouse IP
- Trainable LD50 Mouse IV
- Trainable LD50 Mouse OR
- Trainable LD50 Mouse SC
- Trainable LD50 Rat IP
- Trainable LD50 Rat OR
- hERG Inhibitors
- Trainable hERG I
Calculates probability that the compound will inhibit hERG with IC50 < 10 μM.
- Trainable hERG I
- Ames Test
- Trainable Ames
Calculates probability that the compound will be mutagenic in Ames test.
- Trainable Ames
As a starting point for the calculations a number of Built-in Self-training Libraries with experimental values of the corresponding properties is provided for each ‘Trainable Model’ in ACD/Percepta.
For more information see Trainable Libraries and Training