CYP2C9 Regioselectivity: Difference between revisions
Created page with "==Overview== <br /> The information about possible metabolites of a drug candidate is desirable in the earliest stages of drug development, but the experimental testing of bi..." |
|||
| Line 17: | Line 17: | ||
<br /> | <br /> | ||
[[Image: | [[Image:Cyp2c9_regioselectivity.png|center]] | ||
<br /> | <br /> | ||
Revision as of 13:27, 29 May 2012
Overview
The information about possible metabolites of a drug candidate is desirable in the earliest stages of drug development, but the experimental testing of biotransformations is usually done only in the later phases. The P450 regioselectivity module of ACD/Percepta predicts possible sites of metabolic reactions taking place in human liver microsomes (HLM) and catalyzed by 5 individual cytochrome P450 enzymes (CYP1A2, CYP2C19, CYP2C9, CYP2D6, CYP3A4). The predictive models are based on experimental data for >900 compounds. Metabolites that are formed when incubating the compound with human liver microsomes and recombinant enzymes were considered. Separate probabilistic models were built for 5 different reaction types: aliphatic and aromatic hydroxylation, N-dealkylation, O-dealkylation and S-oxidation.
Six separate regioselectivity modules are provided. "Human Liver Microsomes" module identifies all probable sites of human liver microsomal metabolism in the molecule while the other modules predict regioselectivity of the reactions catalyzed by individual enzymes (CYP1A2, CYP2C19, CYP2C9, CYP2D6, CYP3A4).
Features
- Predicts possibility of being a metabolism site for every atom in the molecule
- Calculates a Reliability Index for every prediction
- Illustrates the predictions in a color-coded manner
- Performs a similarity search and displays top 5 most similar structures from the training set of the model along with their names and experimental results
Interface
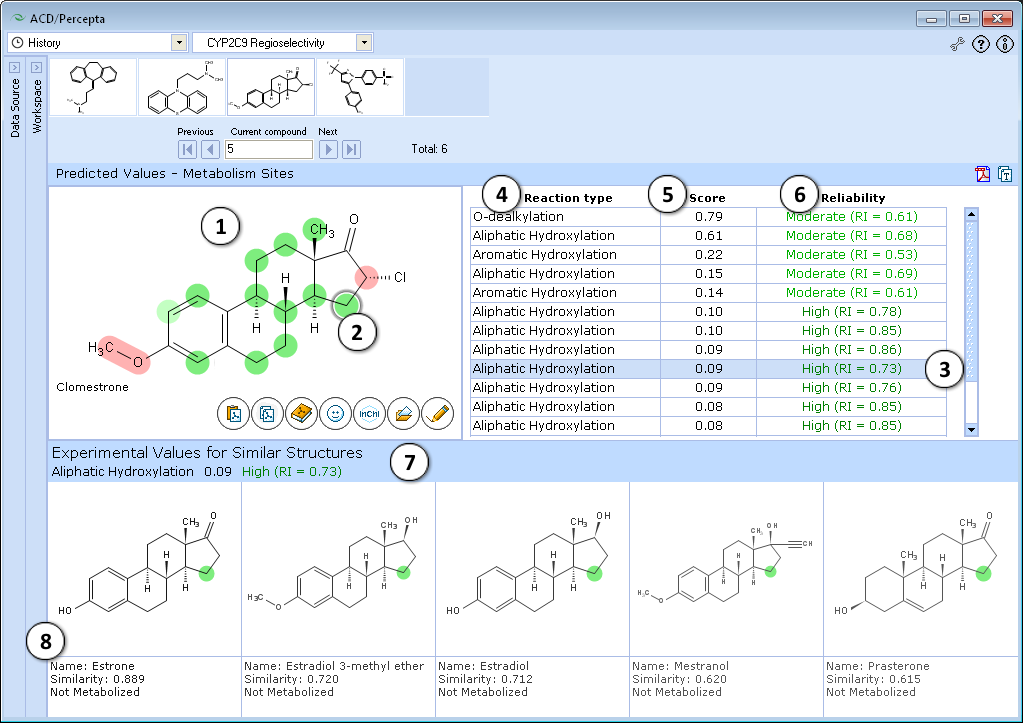
- Predictions are performed for all possible metabolism sites in the molecule and atoms are colored according to predicted values: intensive red corresponds to probability > 0.8; light red – probability in the range 0.6-0.8. Green color indicates non-metabolized atoms (intensive green – probability < 0.2, light green – probability in the range 0.2-0.4), while inconclusive predictions (probability in the range 0.4-0.6) are marked gray.
- Atom with the highest predicted score of metabolism is automatically selected and marked with a circle. Other atoms can be selected by clicking on them or by clicking on the corresponding entry in the table in the right
- Table entry corresponding to currently selected atom is highlighted
- Type of metabolic reaction taking place at the particular site
- Calculated score for an atom to be a metabolism site
- Calculated value of Reliability Index (RI) supporting the prediction
- Prediction for the currently selected atom is outlined on the top of Similar Structures Pane
- Five most similar atoms from the training set are displayed along with experimentally assigned classes and Similarity Indices to the selected atom. Similar atoms are color-marked according to experimental data (red – experimentally determined site of metabolism, green – no metabolism observed, grey - inconclusive data).
Technical information
Predicted endpoints
CYP450 Specificity-related modules in ACD/Percepta provide the following quantitative predictions:
- P450 Substrates: Probability that the compound of interest will be metabolized by a certain CYP450 isoform.
- P450 Inhibitors: Probability that the compound of interest will inhibit a certain CYP450 enzyme with IC50 below defined threshold. Two types of predictive models utilizing different IC50 thresholds have been developed. "General inhibition" models estimate whether the analyzed compound will exhibit any clinically significant CYP450 inhibition at all (IC50 < 50 μM), while "Efficient inhibition" models predict probability that the compound will inhibit selected enzyme with IC50 < 10 μM.
- P450 Regioselectivity: Probability to be metabolized in human liver microsomes (or by a specific CYP450 enzyme) for every atom in the molecule.
Predictions are provided for five major cytochrome P450 isoforms (3A4, 2D6, 2C9, 2C19, 1A2) that are responsible for more than 80% of Phase I metabolism. In addition to Regioselectivity models for individual enzymes, overall HLM Regioselectivity module is also available. This module estimates the overall probabilities of human liver microsomal metabolism taking place at particular sites of the molecule. All predictions are supplied with Reliability Indices (RI) serving as an internal measure of prediction confidence (see Model Features section for more details about RI calculation).
Sources of experimental data
- P450 Metabolism (Substrate specificity and metabolism sites): only experimental data from original scientific publications were used for modeling of cytochrome P450 metabolism sites. The full dataset contained ~700 compounds with >800 possible metabolism sites. The literature dataset was expanded with information about marketed drugs’ metabolism and the expanded dataset was used for cytochrome P450 substrate modeling.
- P450 Inhibition: Two types of experimental data were used for cytochrome P450 inhibition modeling, including data from original scientific publications, information about marketed drugs, as well as data from the NCBI PubChem project.
Data sets
The sizes of the data sets used to develop the predictive models of substrate and inhibitor specificity are presented in the table below:
| Isoform | N (Substrate specificity) | N (Inhibitor specificity, cut-off: IC50 < 50 μM) | N (Efficient inhibition, cut-off: IC50 < 10 μM) |
|---|---|---|---|
| CYP1A2 | 935 | 4867 | 5815 |
| CYP2C9 | 867 | 7666 | 7677 |
| CYP2C19 | 794 | 6899 | 6833 |
| CYP2D6 | 1001 | 7707 | 7507 |
| CYP3A4 | 960 | 6684 | 7927 |
Regioselectivity models were based on experimental data for 873 compounds collected from publications dealing with analytical identification of the metabolites observed after the incubation of compound with human liver microsomes or recombinant cytochrome P450 enzymes. Every carbon atom with at least one hydrogen attached was marked as a site of metabolism, if hydroxylation at the atom was observed, or site of no metabolism otherwise. For dealkylation reactions, carbon atoms of the leaving groups were marked in the same manner. Some sites were marked as "inconclusive" and consequently not used in the modeling. The table below shows the overall number of marked atoms used for building the models:
| No. of atoms | HLM | CYP3A4 | CYP2D6 | CYP2C9 | CYP2C19 | CYP1A2 |
|---|---|---|---|---|---|---|
| Positive | 1269 | 795 | 354 | 288 | 249 | 383 |
| Inconclusive | 340 | 176 | 49 | 43 | 15 | 61 |
| Negative | 7182 | 6757 | 6305 | 6314 | 5210 | 6020 |
| Total | 8791 | 7728 | 6708 | 6645 | 5474 | 6464 |
Fully searchable CYP450 Specificity databases are not available in the current version of ACD/Percepta, yet each prediction performed by P450 Substrates and Inhibitors modules is displayed along with experimental data for five compounds from the training set most similar to the molecule of interest. The provided information for similar compounds includes classification as substrates/non-substrates (inhibitors/non-inhibitors at two IC50 cut-offs) of the relevant CYP450 enzyme assigned on the basis of experimental results together with original references. In case of Regioselectivity predictions five most similar atoms from the training set are shown with color-marks indicating whether a metabolic reaction taking place at the particular site of the molecules was observed experimentally.
Model features & prediction accuracy
The model was developed with Algorithm Builder using a novel methodology consisting of two parts:
- Global baseline statistical model employing binomial PLS with multiple bootstrapping using a predefined set of fragmental descriptors.
- Local correction to baseline prediction based on analysis of experimental data for similar compounds.
The underlying methodology enables obtaining an intrinsic evaluation of prediction confidence by the means of Reliability Index (RI) values calculated for each prediction. RI ranging from 0 to 1 serves as an indication whether a submitted compound falls within the Model Applicability Domain. Two criteria influence the calculation of Reliability Index of a prediction:
- Similarity of the analyzed molecule to compounds in the Self-training Library (prediction is unreliable if no similar compounds have been found in the Library).
- Consistency of experimental data for similar compounds (discrepant data for similar molecules lead to lower RI values).
The predictive models of CYP450 substrate and inhibitor specificity are also Trainable meaning that their Applicability Domains may be expanded to account for the ‘in-house’ experimental data available in your company without the need to rebuild the baseline statistical model from scratch. Addition of new compounds to the module Self-training Library results in an instant improvement of prediction accuracy for the respective compound classes. Moreover, addition of 'in-house' data allows adapting the existing model to the particular experimental protocol used in your company and avoiding potential issues related to discrepancies between different experimental methods used for determination of drug interactions with CYP450 enzymes.
If the compound is within model Applicability Domain (acceptable reliability index) accuracy and sensitivity of classification is close to 90% for inhibitors and close to 80% for substrates. The accuracy of in silico prediction of cytochrome P450 inhibitors is comparable to the screening results. Metabolism sites are predicted correctly if there are similar metabolism sites in the dataset and if the reliability of prediction (RI value) is high.